Comparative transcriptome analysis and phenotypic monitoring of Trifolium pratense (Fabaceae) under land use scenarios
TRATSCH uses comparative transcriptome analysis and phenotypic monitoring of T. pratense (Fabaceae) under land use scenarios to investigate the causes of phenotypic plasticity in response to mowing/grazing.

The most important impacts on existing structures of biodiversity and ecosystem processes are caused by changes in land use. However, knowledge on how organisms cope with environmental perturbations and changes in land use on the molecular level is scarce.
Here we investigate differential gene expression and changes in morphology of the agronomical important non-model plant species Trifolium pratense at different sites and treatments. Further, differentially expressed growth regulators will be analyzed by transgenic approaches to elucidate their role in phenotypic plasticity.
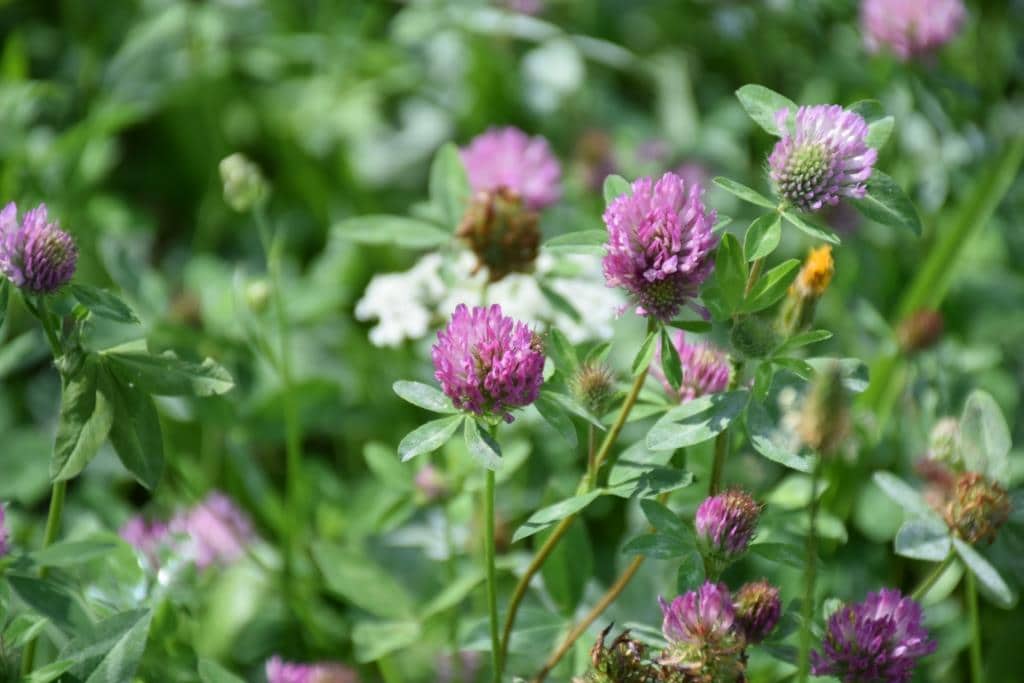
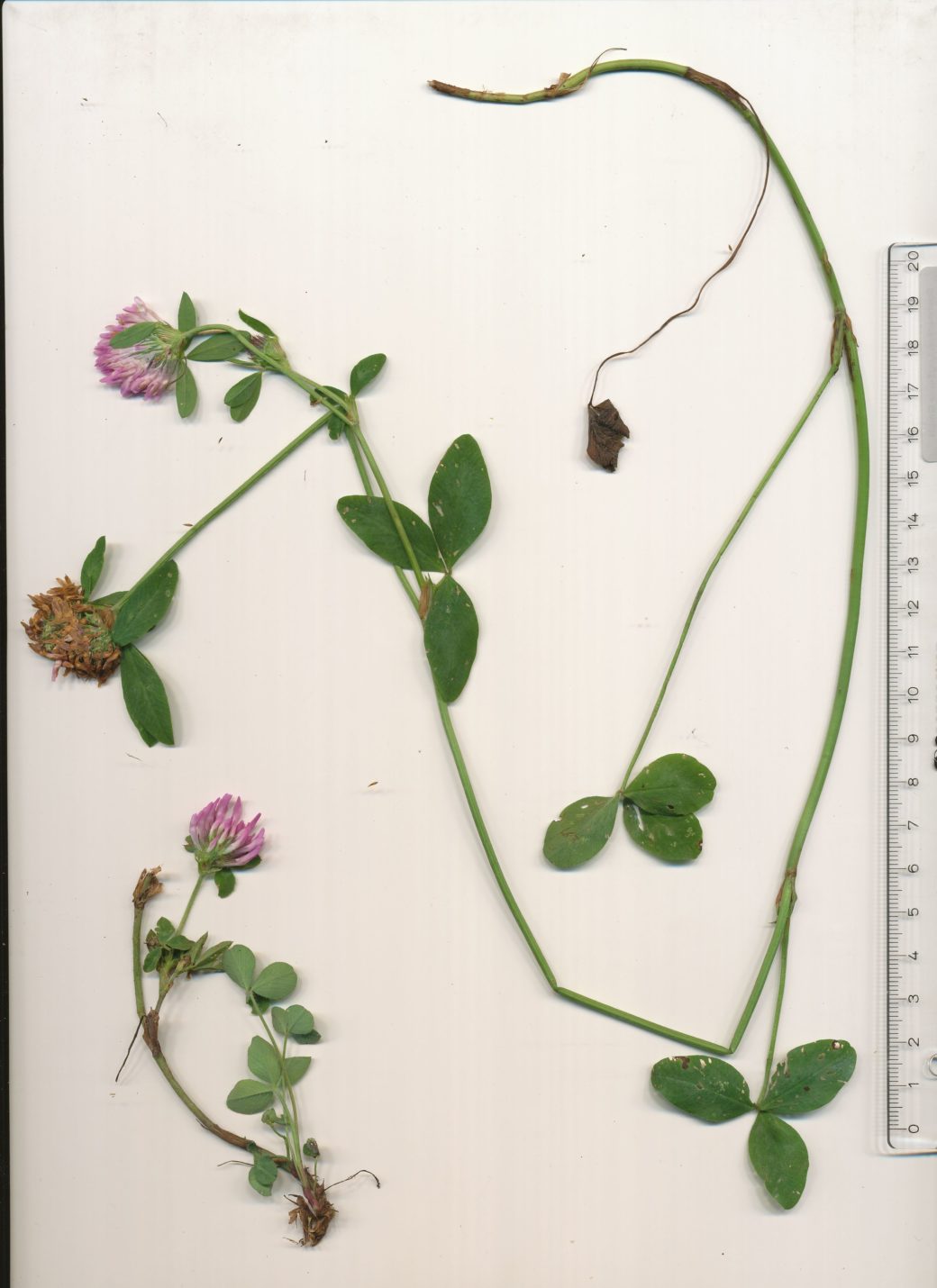
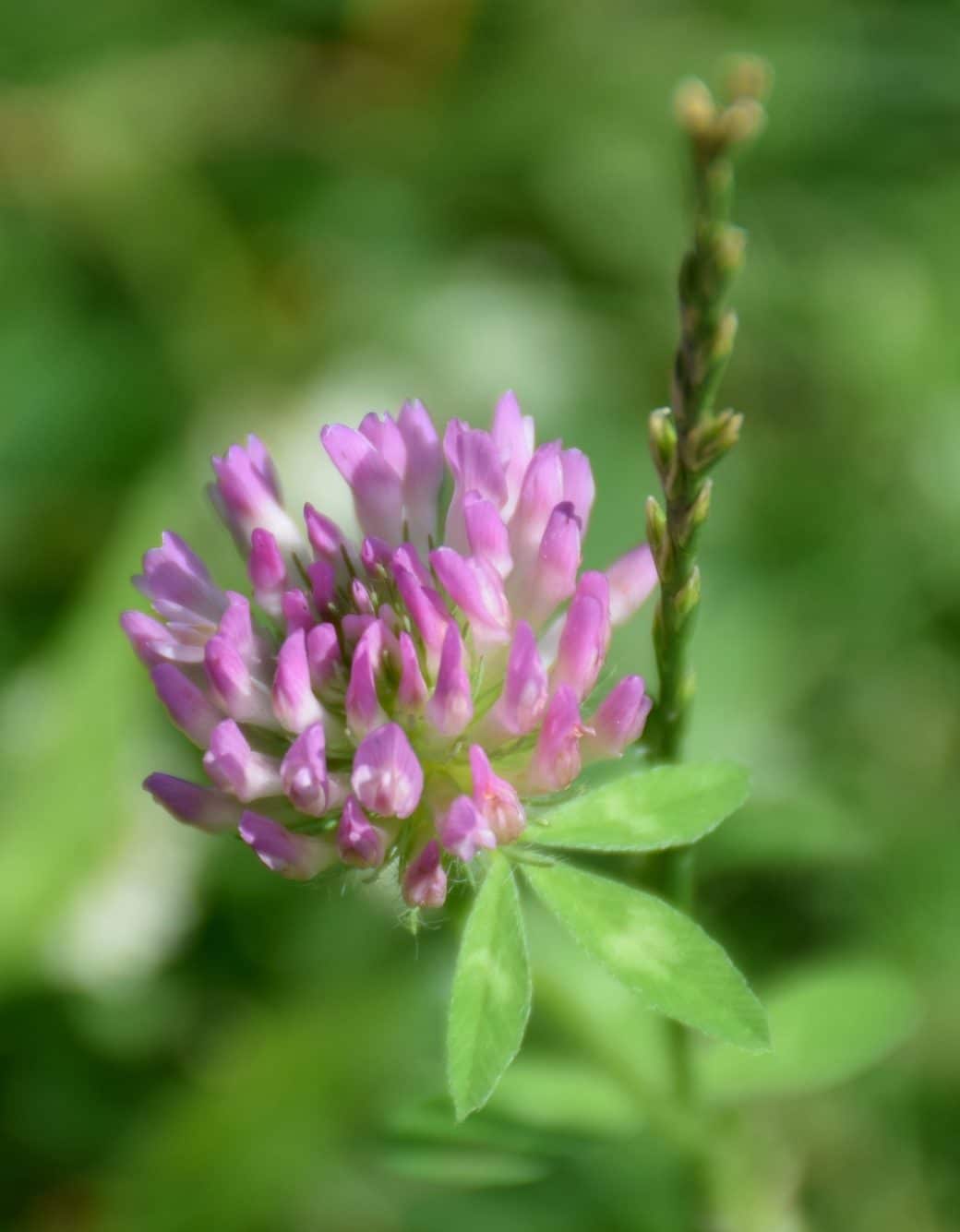
As a common grass-land species T. pratense is present on most grassland sites of the Biodiversity Exploratories. Furthermore T. pratense shows drastic morphological changes upon grazing/mowing. While the undisturbed plants grow up to 70 cm in height producing mainly a single shoot, the grazed-on plants are dwarf, produce more side shoots and smaller, rounder leaves on shorter petioles.
In a collaborative effort we investigate the molecular nature of morphological differences of T. pratense at different sites under different land use scenarios and treatments, by applying transcriptome analysis in combination with functional analysis of developmental regulators, transcriptome fingerprinting techniques and phenotypic monitoring.
a) Red clover (Trifolium pratense) shows drastic phenotypic plasticity in response to mowing/grazing.
b) Differentially expressed genes regulating growth and development are responsible for morphological changes.
c) Changes in gene expression to mowing/grazing of T. pratense are species specific and independent of biotic and abiotic factors.
1) Exploring the phenotypic appearance of T. pratense.
2) Investigating the intra-population and intra-specific genetic response variability across different land use scenarios by transcriptome fingerprinting (cDNA-ddRADseq).
3) Analysis of transcriptomes generated from T. pratense plants to detect differentially expressed genes.
4) In-depth expression analysis of candidate genes for morphological changes by qRT-PCR.
5) Gene knock-down approaches for functional analysis of candidate genes.
6) Data correlation between sub-projects, sites, and site specific environmental data.
Quantitative next generation sequencing cDNA fingerprinting techniques (cDNA-ddRADseq) and phenotypic monitoring will be applied on repetitive site and treatment specific individuals across different land use scenarios and correlated to environmental facts. The results will provide large numbers of comparative characters, sampled on many individuals across many sites, which will allow for pattern identification of treatment specific and/or environment related plastic plant responses to perturbations.
Transcriptome profiling will be achieved on a limited number of individuals and sites with deeper insights on populations´ gene expression. RNAseq allows for monitoring changes in transcriptome composition between populations with divergent treatments/phenotypes. The transcriptome analysis will provide candidate genes for further functional analysis, which will lead to an understanding of the underlying molecular mechanism governing complex changes to phenotypes resulting in phenotypic plasticity in response to different treatments across the individual sites.
By combining our results with other data generated on the sites of the Biodiversity Exploratories (e.g. soil parameter, climate data) we aim 1) to identify genes and gene functions that matter most in a given ecological interaction and 2) to untangle the simultaneous and interacting effects of land use scenarios on T. pratense.