BELongDead – Fungal Functional Diversity – Halle/Zittau/Grafenau II – Structural and functional diversity of fungi and bacteria in deadwood
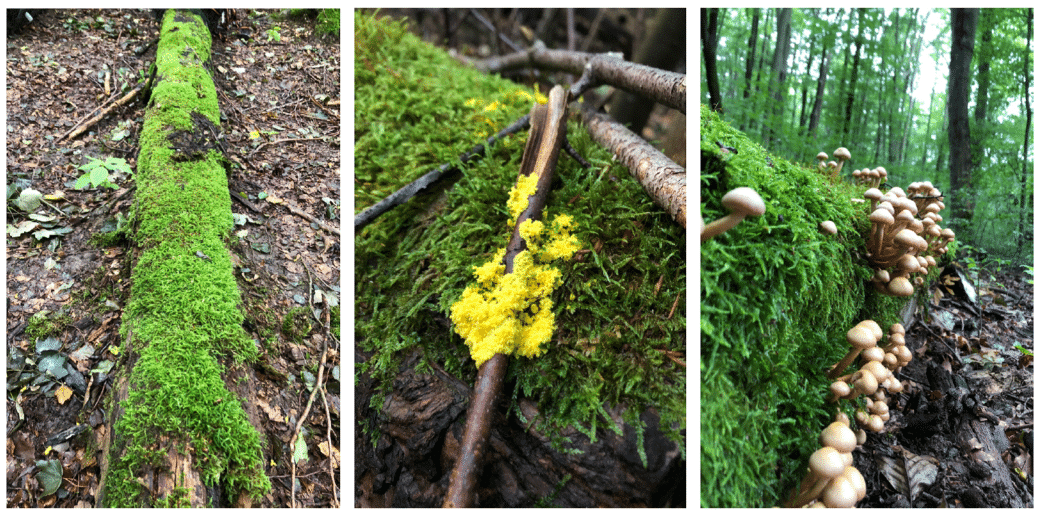
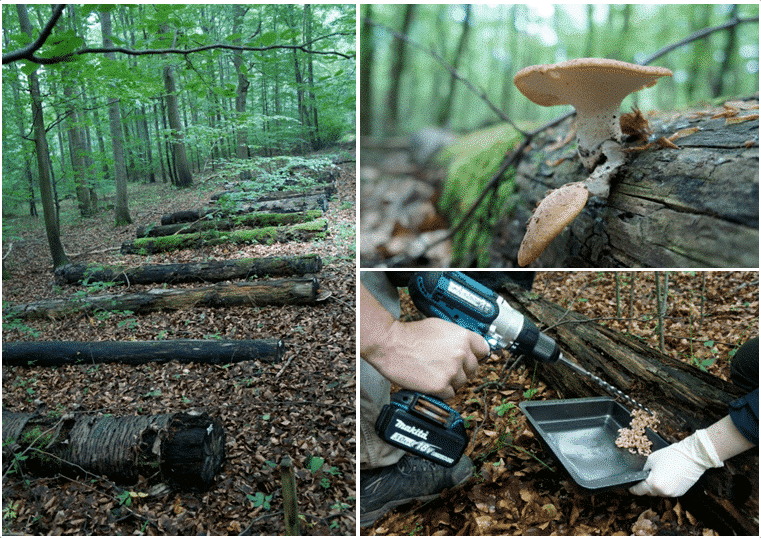
Compared to the previous projects FunWood I-II, which was largely concerned with the influence of forest management on microbial diversity in existing decomposing deadwood under field conditions, we are now shifting our focus to an experimental platform. Das BELongDead experiment was initiated in 2008 under the leadership of Prof. Dr. E.D. Schulze (MPI Biogeochemistry Jena) with the aim of investigating the influence of the surrounding habitat on deadwood and its decomposition processes. Another focus is on the long-term observation of the colonisation of the deadwood logs by a wide variety of organisms.
The aim is to investigate to what extent I) the surrounding ecosystem influences deadwood decomposition, II) how deadwood colonisation occurs and III) how microorganisms control deadwood decomposition and thus influence ecosystem processes such as nutrient turnover. BELongDead allows us to study the influence of land use in the form of forest management on a standardised set of deadwood (13 different tree species of the same size and decomposition onset), evenly distributed in 3 replicates in the 3 exploratories and in 3×3 differently managed plots each.
The aim of our project is to combine state-of-the-art molecular biological methods with classical fruiting body mapping and spore collections to I) determine the type and quantity of wood decomposition and different forest management aspects, II) observe and study distribution and succession patterns of fungi over time, III) determine fungal activity at the transcriptome and enzyme levels and correlate these results with process data, IV) determine resulting changes in wood chemistry, V) determine the influence of functionally different bacteria on fungal diversity, and VI) ultimately identify key species in these complex processes.
Meanwhile, the experiment reaches medium decomposition levels in different tree species. The following central hypotheses were derived:
1. increased forest management intensity reduces the species pool of wood-inhabiting fungi at the landscape and forest stand level.
2. intensive forest management is a habitat filter that favours certain species with certain life strategies (e.g. generalists).
3. forest management relaxes competitive interactions between wood-dwelling fungi, leading to higher wood decomposition rates.
4. wood decomposition processes are highly predictable from fruiting body dating combined with molecular data on phylogenetic and functional diversity.
5. there are very high bacterial diversities and certain bacteria occur non-randomly with certain fungi.
6. white rot fungi are the most relevant fungi in deadwood degradation and primarily use manganese peroxidases to degrade lignin and increase element bioavailability.
To answer our questions, we use state-of-the-art molecular biology techniques at the DNA and RNA level, including so-called “Next Generation Sequencing” techniques (NGS). Wood chemical parameters and enzymes are also determined using current methods. In addition to intensive fruiting body mapping, spore collectors are installed to identify airflow-based distribution patterns and, on the other hand, to be able to compare which fungi are potentially present (species pool) and which are already established in deadwood.