Drivers and functions of foliar endophyte communities
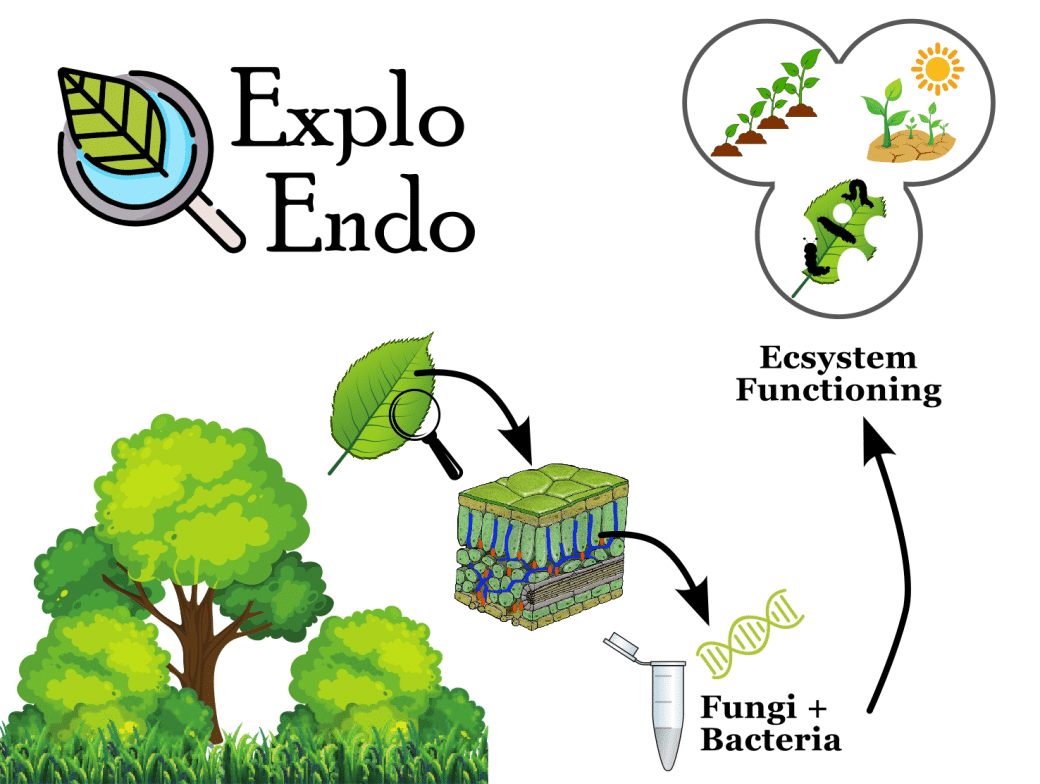
Foliar epiphytes are ubiquitous microscopic fungi and bacteria that inhabit the leaf tissue of plants without causing any apparent harm. Instead, foliar endophytes can promote plant growth and nutrient cycling, and mitigate stress from drought, pathogens and herbivores. Yet, beside a growing scientific interest, there are still significant gaps in our understanding of the distribution, diversity, drivers and ecological roles of foliar endophyte communities, especially with regard to potentially interacting effects of land-use intensity and plant diversity .
We aim at investigating the diversity, drivers, and ecological associations of foliar endophyte communities with two approaches:
- (1) host plant-specific analyses of foliar endophytes DNA in 15 target plant species in forests and grasslands and
- (2) environmental DNA-analyses in time series of dried grassland biomass.
(H1) Within target host-plant species, the diversity of foliar endophyte communities is positively related to the diversity of co-occurring plants and regional habitat types and negatively related to increasing land-use intensity.
(H2) Due to differences in species richness and physical barriers, we expect relationships between plant diversity and foliar endophytes to vary between grassland and forests habitats.
(H3) Summarised across grassland biomass, we expect a reduction in plant diversity, an increase in land-use intensity, and an increase in drought stress to all be related with a higher inter-annual turnover of leaf-associated fungal communities.
For the host plant-specific approach, we will conduct an extensive field sampling in all experimental grassland and forests plots to collect leaf samples from the following common 15 target plant species:
(1) Fagus sylvatica L. , (2) Acer pseudoplatanus L., (3) Picea abies (L.) H. Karst., (4) Pinus sylvestris L., (5) Milium effusum L., (6) Carex sylvatica Huds., (7) Anemone nemorosa L., (8) Urtica dioica L., (9) Poa trivialis L., (10) Dactylis glomerata L., (11) Cerastium holosteoides Baumg., (12) Trifolium repens L., (13) Veronica chamaedrys L., (14) Plantago lanceolata L., (15) Alopecurus pratensis L.
All leaf samples will be stored on ice and transported to the laboratories of the Martin Luther University Halle-Wittenberg, where we sterilize the surface and then homogenize each leaf sample.
For the environmental DNA-approach, we will collaborate with Core Project 5 to analyse dried and homogenized samples of grassland aboveground biomass from 2018-2025.
From all biomass samples, we will extract the DNA a gDNA Plant Kit (Invitrogen™, ChargeSwitch®, Thermo Fisher, Munich, Germany) and amplify/purify the Internal Transcribed Spacer 1 gene region for fungi (using primers ITS1F and ITS2) and the V4 region of the 16S rRNA gene for bacteria (using primers F515 and R806). After equimolar pooling, 380 combined fungi and bacterial DNA-samples (to minimize costs) will be Illumina-sequenced. The obtained DNA reads will be processed with a DADA2 pipeline and assigned to fungal and bacterial amplicon sequence variance, whose distribution and diversity we will analyse with generalized mixed-effects, boosted regression, and multivariate models.