Epigenetic diversity of grassland plants in the Biodiversity Exploratories
The biological diversity within species is an important component of biodiversity. It is the raw material for evolution and adaptation, and it can be an important driver of population and ecosystem processes such as productivity, stability or invasion resistance.

In the past, within-species diversity was assumed to reflect genetic (= DNA sequence) diversity. However, recent evidence suggests that, even in the absence of genetic diversity, within-species variation in ecologically important traits can also be created by underlying epigenetic variation. Epigenetic modifications of the genome, such as DNA methylation or various histone modifications, which influence gene expression and thus the phenotypes of organisms, can vary across individuals or populations of the same species, and this variation can be heritable.
One main finding of the previous projects QuantGen, GenLink and GenLand is that several common grassland plant species are strongly phenotypically differentiated in relation to land use but show very little corresponding differentiation at the level of DNA sequence. It is an intriguing question whether these species are perhaps differentiated at the epigenetic level, and if such epigenetic differences are related to phenotypic and habitat differences.
In our project we study the extent and ecological significance of epigenetic diversity in grassland plants. Specifically, we:
- quantify epigenetic diversity within and among populations of common grassland plants.
- test the effects of land use on epigenetic diversity.
- examine relationships between plant genetic, epigenetic and phenotypic diversity.
- build expertise in established and next-generation molecular methods for ecological-epigenetic studies.
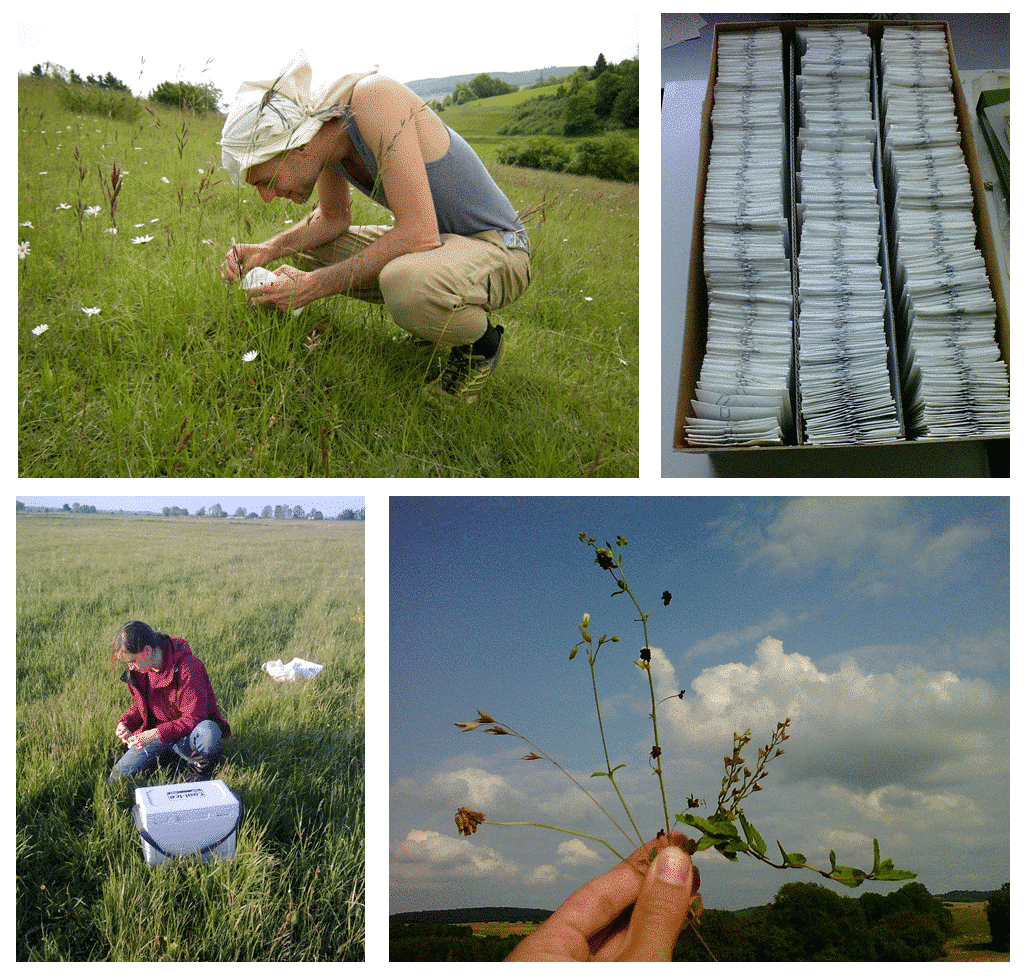
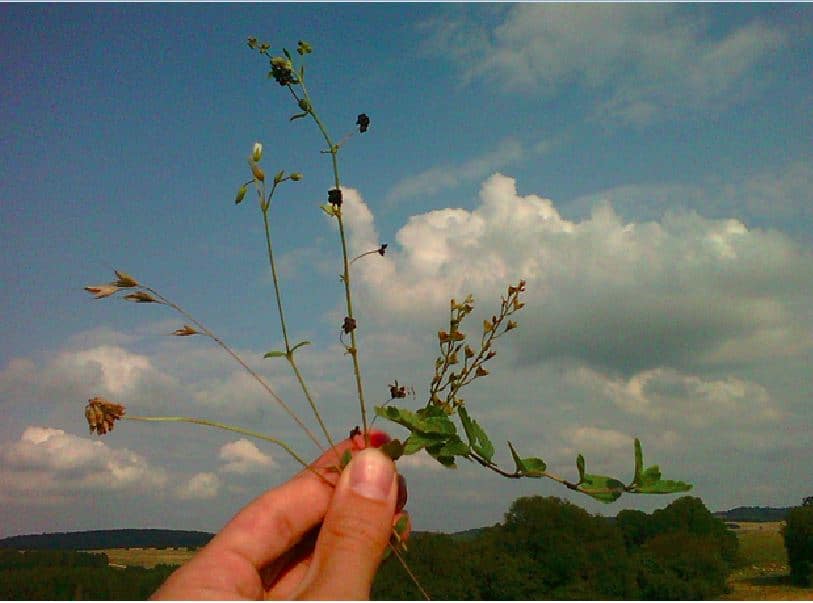
We will study 3-5 of the following grassland species: Bromus hordeaceus, Cerastium holosteoides, Trifolium repens, (Medicago lupulina, Veronica chamaedrys), with 36 populations of each species across the three exploratories. We collect leaf samples in the field and use methylation-sensitive AFLP (MS-AFLP or MSAP) to analyse natural epigenetic diversity – polymorphisms in DNA-methylation at multiple neutral loci across the whole genome. In addition, we will grow a subset of the species from field-collected seeds in a common garden and also analyse their epigenetic diversity with MS-AFLP, in order to determine the heritability of epigenetic variation. Finally, we will analyse a subset of the samples through next-generation sequencing of bisulfite-converted DNA, a novel method of rapid and high-resolution epigenotyping that can identify DNA methylation polymorphisms at the single-nucleotide level (single methylation polymorphisms, SMPs).